graph TD
H123["$$H_0^{(123)}: \left( \theta_1, \theta_2, \theta_3 \right) = \left( \theta_1^0, \theta_2^0, \theta_3^0 \right)$$"]
H12["$$H_0^{(12)}: \left( \theta_1, \theta_2 \right) = \left( \theta_1^0, \theta_2^0 \right)$$"]
H13["$$H_0^{(13)}: \left( \theta_1, \theta_3 \right) = \left( \theta_1^0, \theta_3^0 \right)$$"]
H23["$$H_0^{(23)}: \left( \theta_2, \theta_3 \right) = \left( \theta_2^0, \theta_3^0 \right)$$"]
H1["$$H_0^{(1)}: \theta_1 = \theta_1^0$$"]
H2["$$H_0^{(2)}: \theta_2 = \theta_2^0$$"]
H3["$$H_0^{(3)}: \theta_3 = \theta_3^0$$"]
H123 --> H12
H123 --> H13
H123 --> H23
H12 --> H1
H12 --> H2
H13 --> H1
H13 --> H3
H23 --> H2
H23 --> H3
style H123 fill:#4a90d9,color:#fff
style H12 fill:#7ab3e0,color:#fff
style H13 fill:#7ab3e0,color:#fff
style H23 fill:#7ab3e0,color:#fff
style H1 fill:#a8d1f0,color:#333
style H2 fill:#a8d1f0,color:#333
style H3 fill:#a8d1f0,color:#333
Domain selection and familywise error rate for functional data: A unified framework
Department of Mathematics Jean Leray, UMR CNRS 6629, Nantes University, Nantes, France
March 20, 2026
Team members
| K. Abramowicz, S. Sjöstedt de Luna | Dept. of Mathematics and Mathematical Statistics, Umeå University, Sweden |
| L. Schelin | Dept. of Statistics, Umeå School of Business, Economics and Statistics, Umeå University, Sweden |
| A. Pini | Dept. of Statistical Sciences, Università Cattolica del Sacro Cuore, Milan, Italy |
| S. Vantini | MOX, Dept. of Mathematics, Politecnico di Milano, Milan, Italy |
Context
Problem formulation
Functional data on \(L^2(D) \cup \mathcal{C}^0(D)\), where \(D \subset \mathbb{R}^d\).
A null hypothesis \(H_0^t\) which is true on a subset \(D_0 \subseteq D\) and false on \(D_1 = D \setminus D_0\).
Aim: estimate \(D_0\) by testing locally \(H_0^t\) against \(H_1^t\).
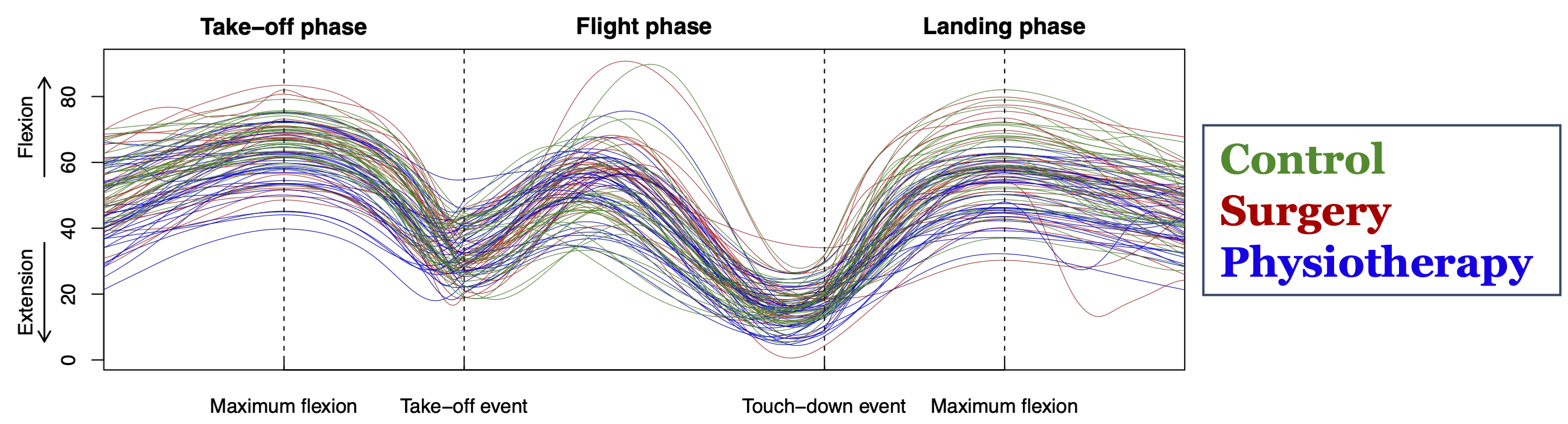
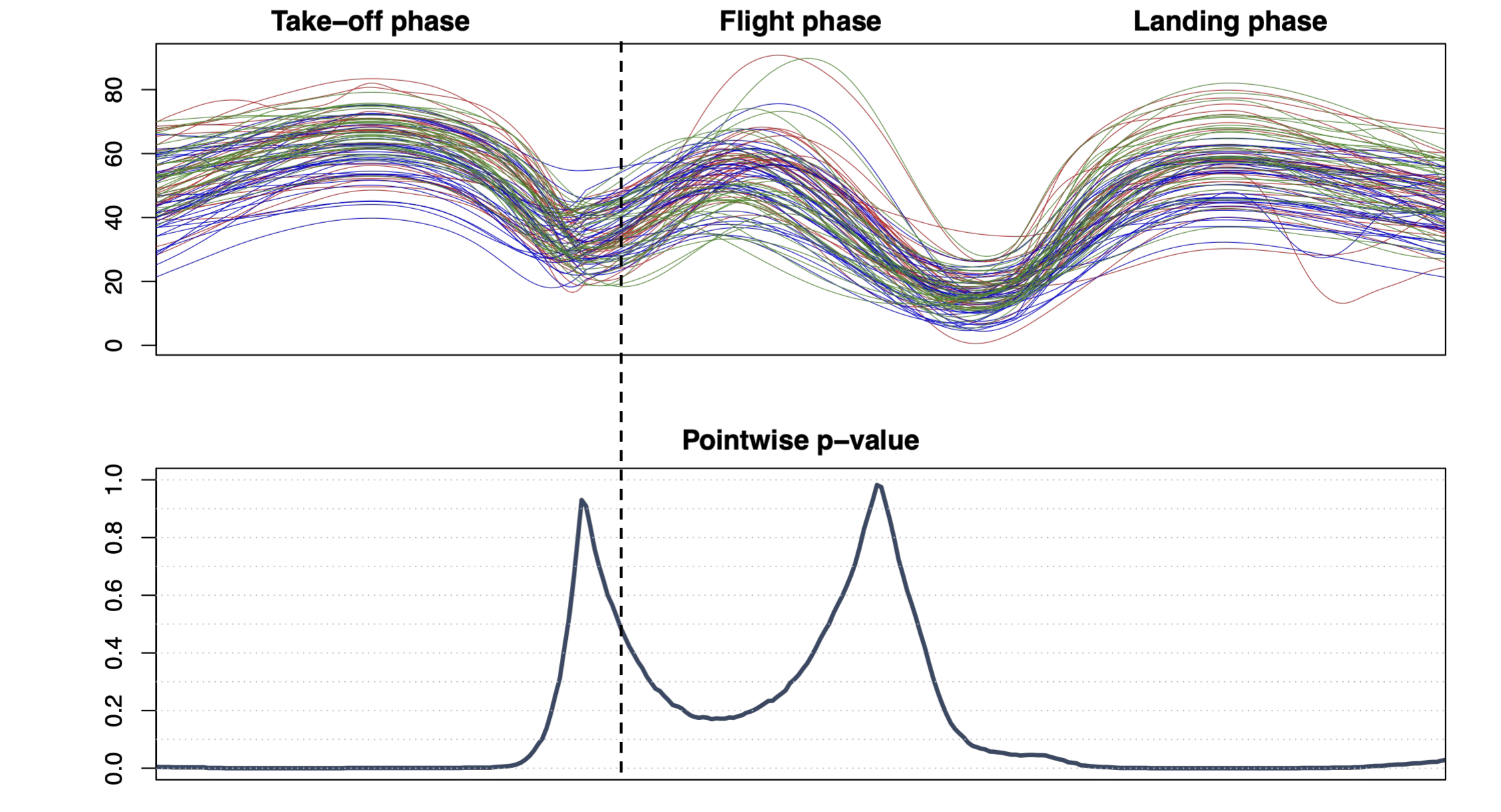
\[ \small{ y_i(t) = \beta_0(t) + \beta_\mathrm{Jump}(t) x_{\mathrm{Jump},i} + \beta_\mathrm{R}(t) x_{\mathrm{R},i} + \beta_\mathrm{PT}(t) x_{\mathrm{PT},i} + \varepsilon_i(t), \quad i = 1, \dots, 95 } \]
\[ \small{ H_0: \beta_\mathrm{PT}(t) = 0 \quad \text{for all } t \in D \quad \text{vs.} \quad H_1: \beta_\mathrm{PT}(t) \neq 0 \quad \text{for some } t \in D } \]
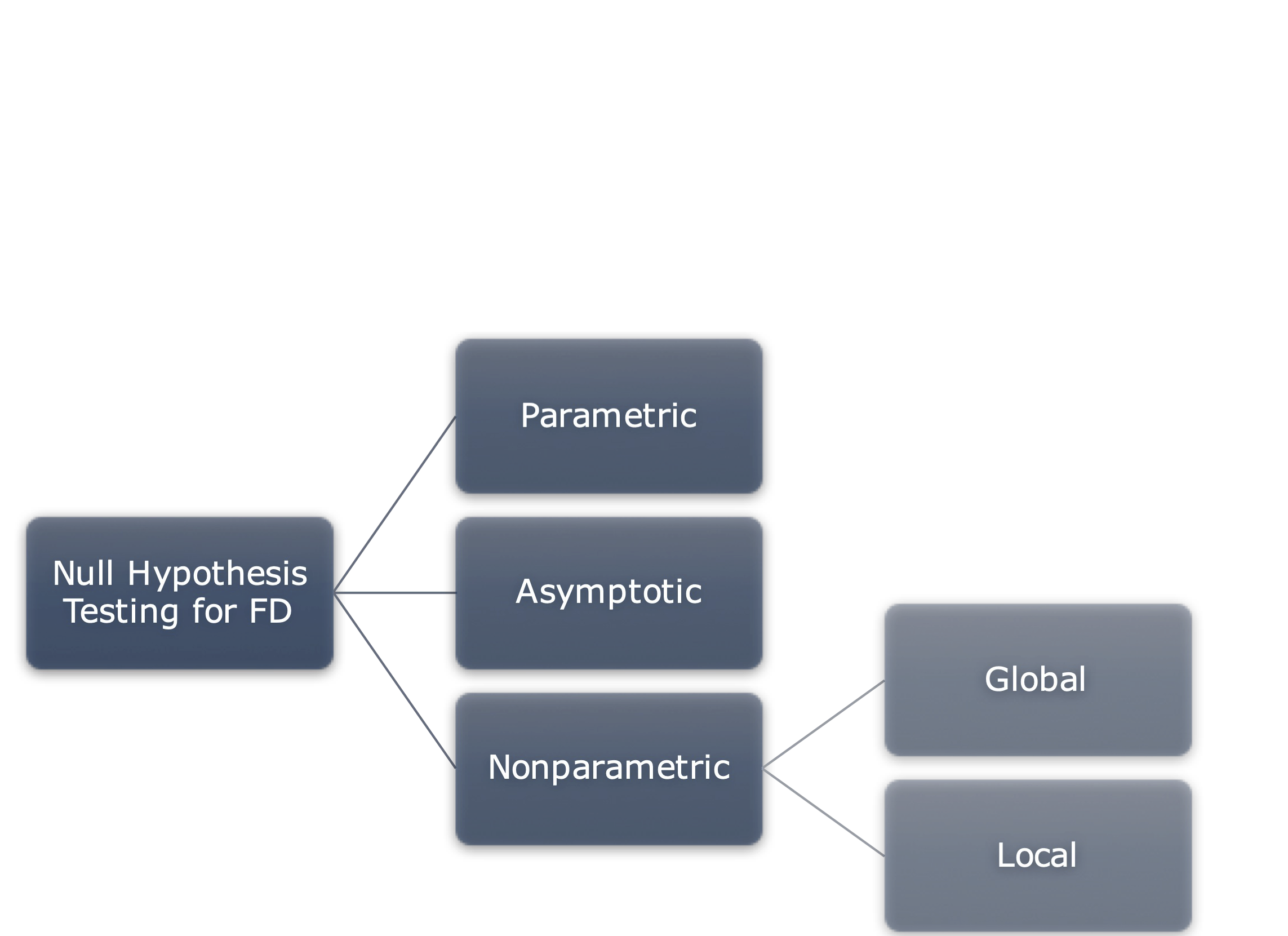
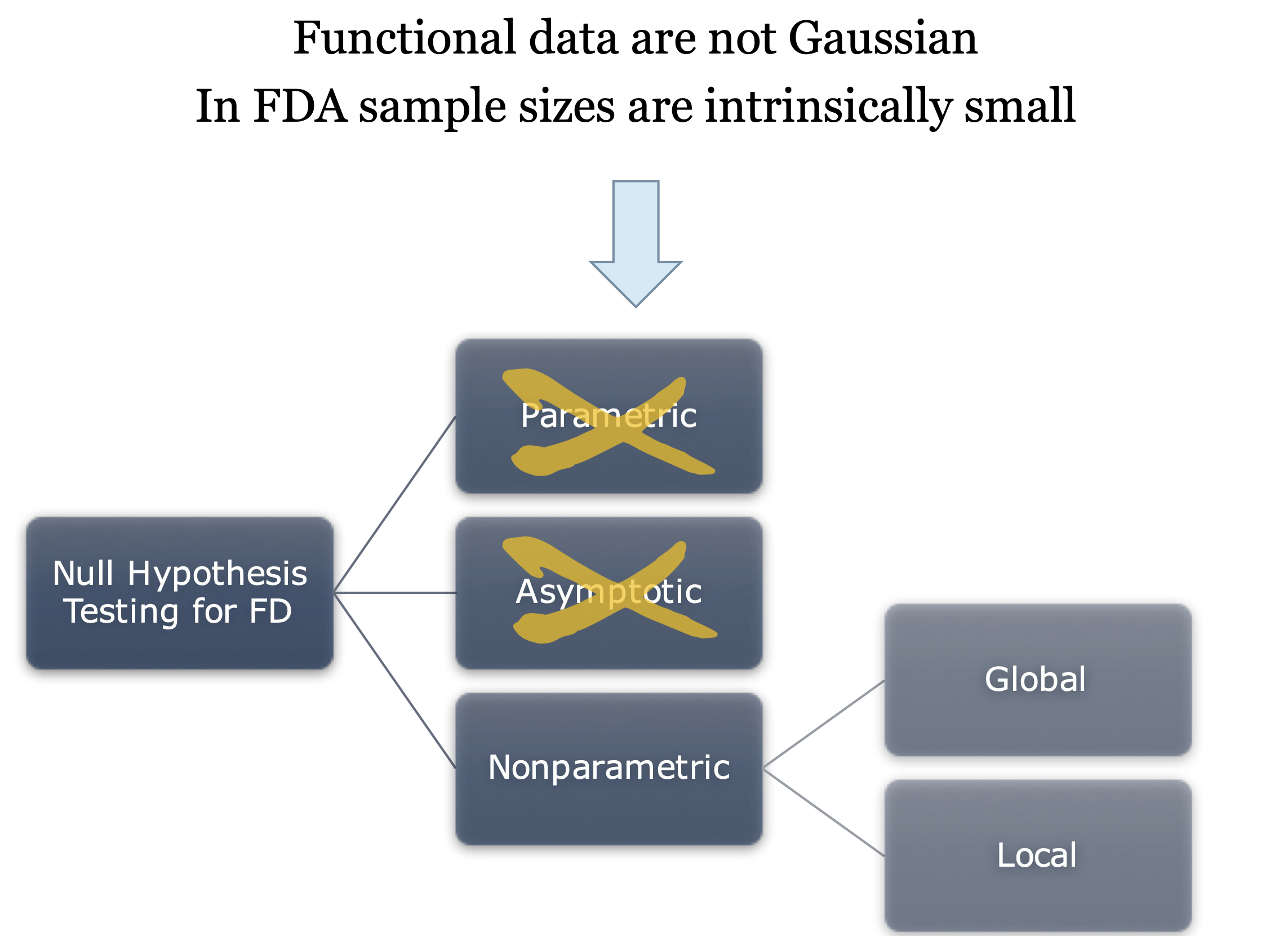
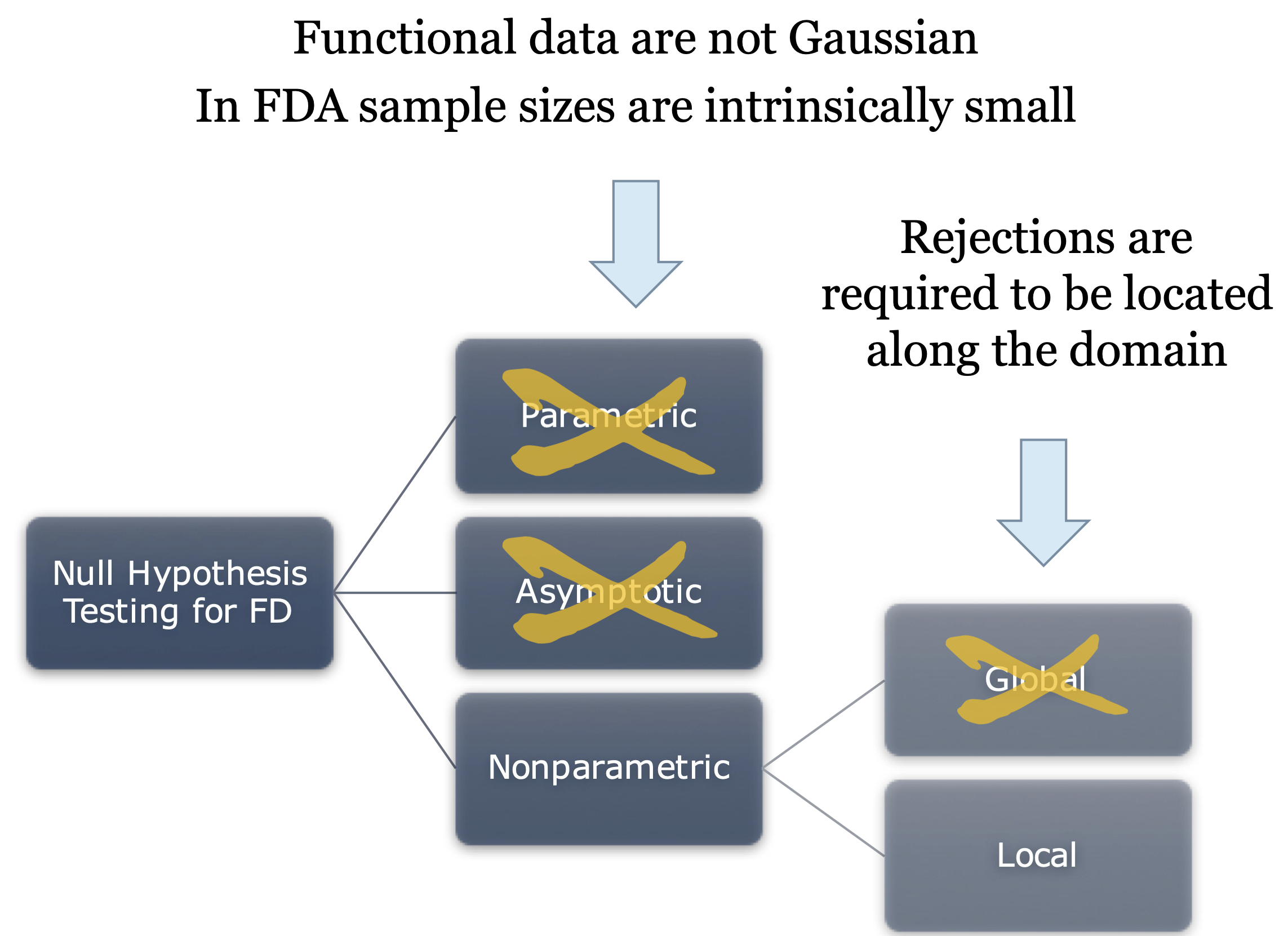
Null hypothesis testing in FDA
Pointwise testing
Straightforward solution: a statistical test for each point of the domain
What is the multiplicity issue?
Definition 1 (Familywise error rate)
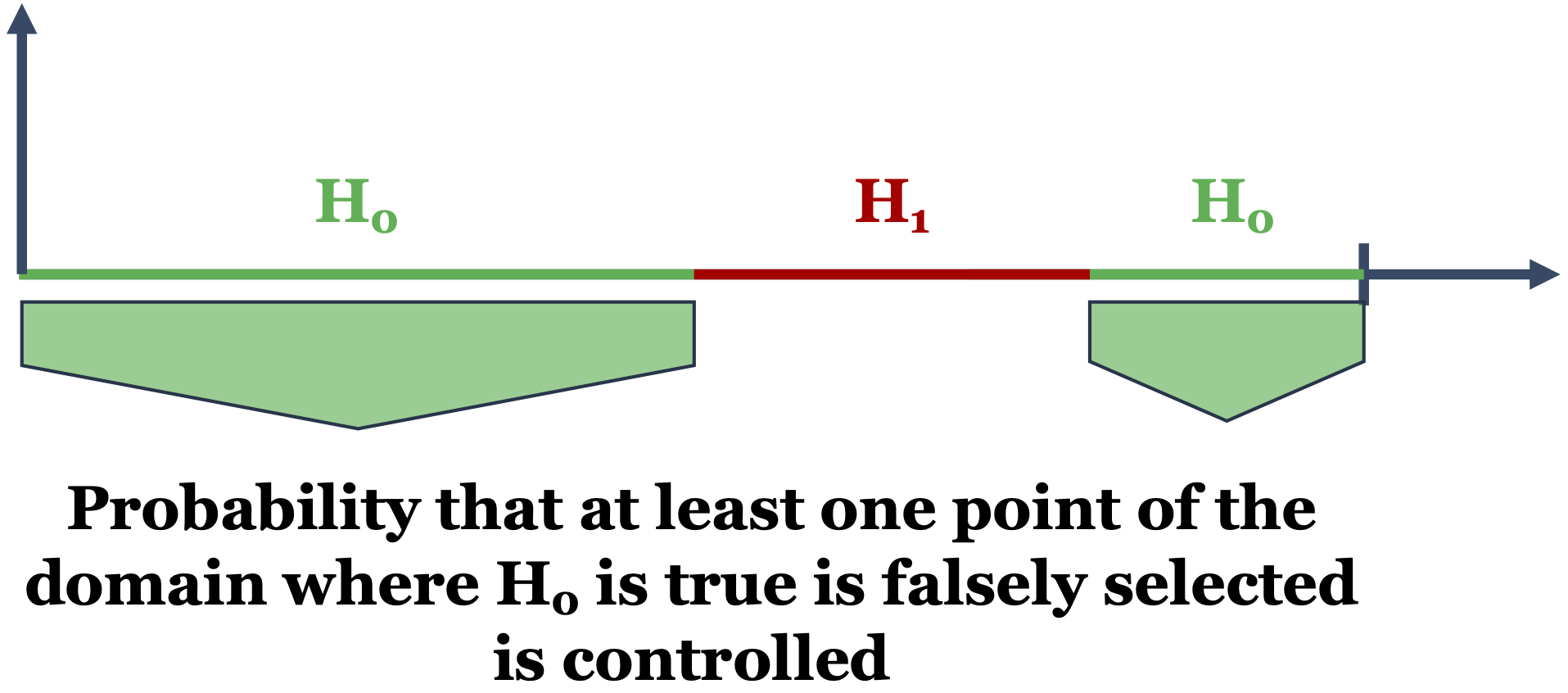
The familywise error rate (FWER) is the probability of making at least one false rejection among a family of tests.
Problem
For \(m\) stochastically independent tests at level \(\alpha\), among which \(m_0\) are true null hypotheses, the FWER can be expressed as: \[ \text{FWER} = 1 - (1 - \alpha)^{m_0} \to 1 \quad \text{as } m_0 \to \infty \text{ when } \alpha \in (0,1]. \]
Unified framework for multiple testing in FDA
Functional control of the FWER
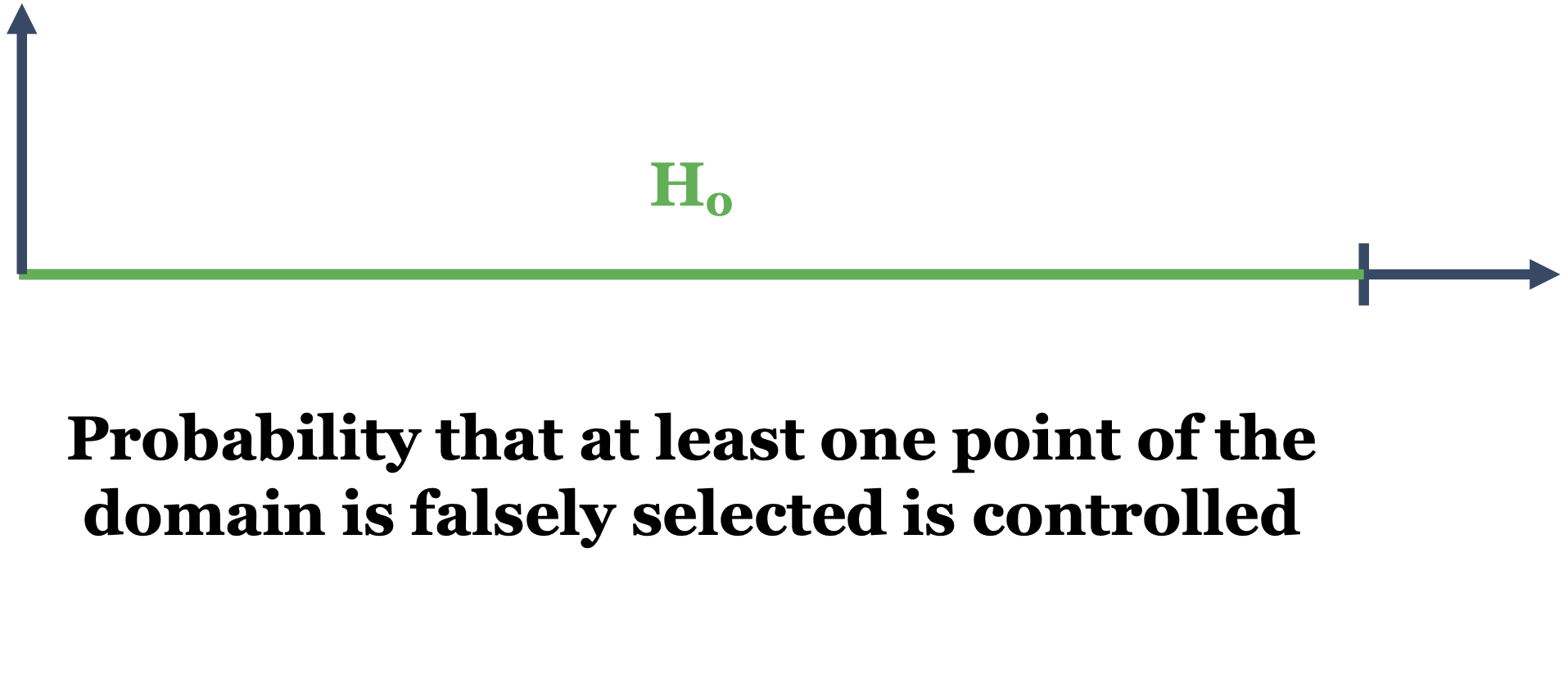
Definition 2 (Weak control of the FWER)
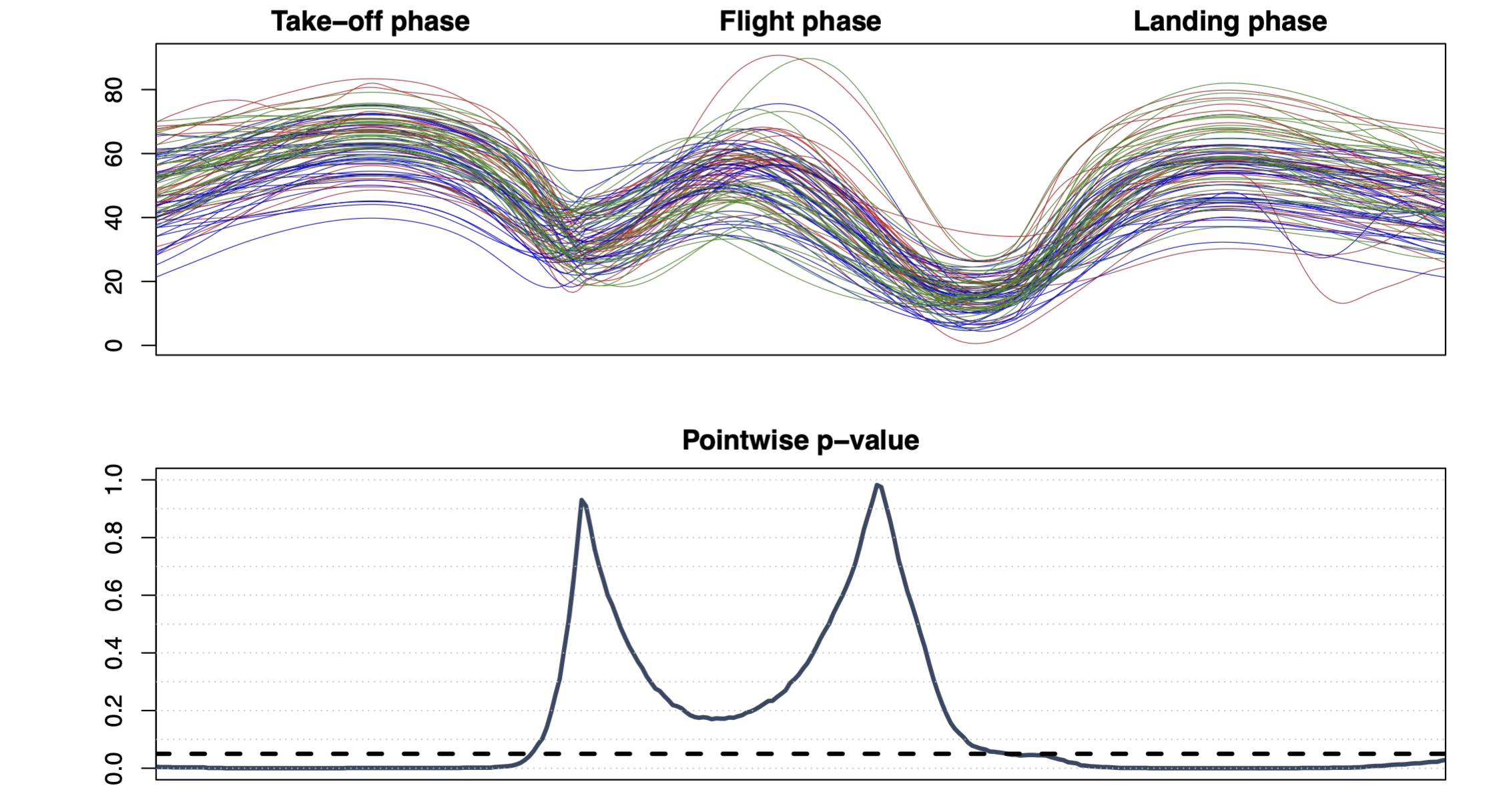
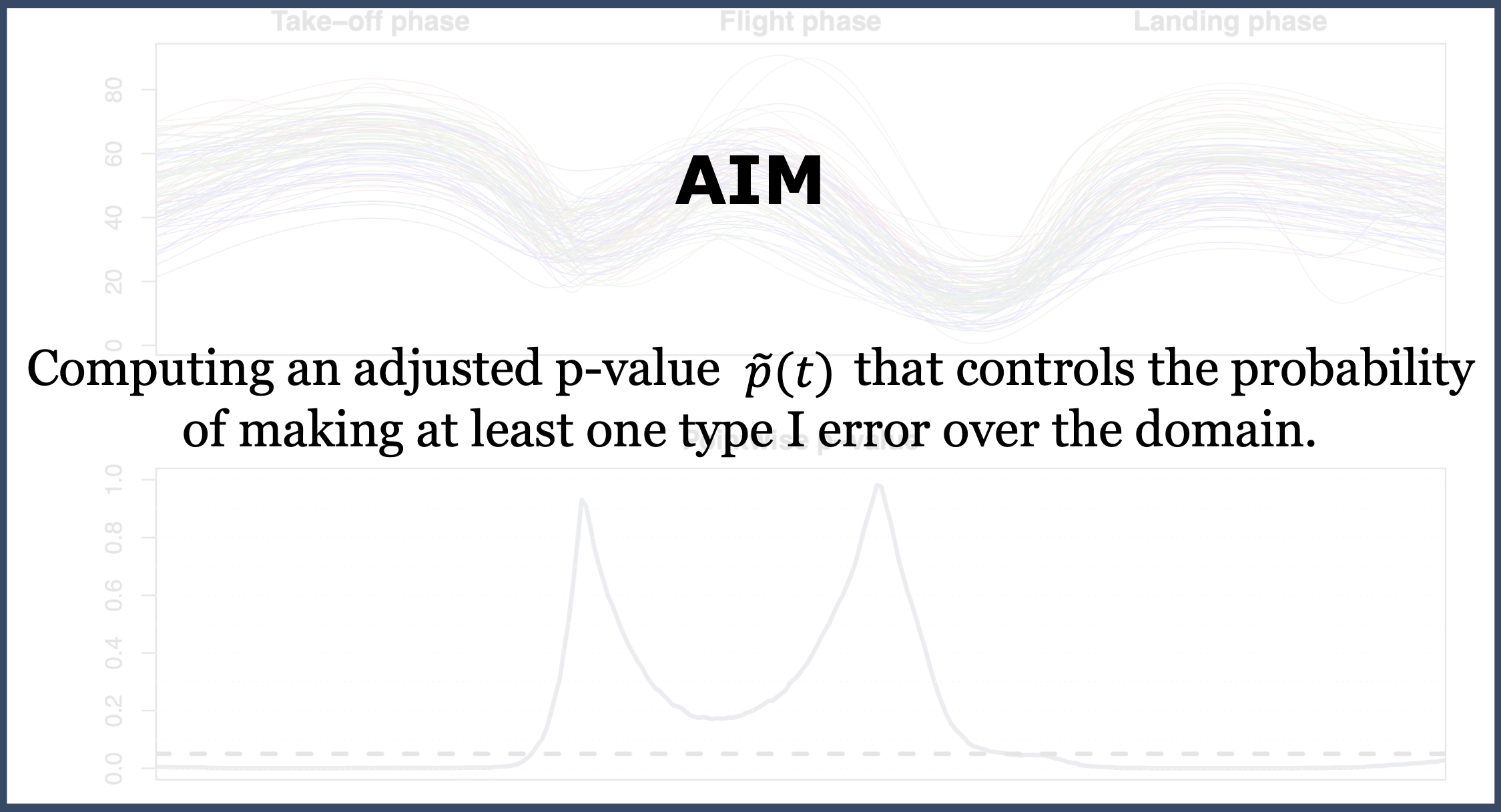
A procedure for locally testing – over the domain \(D\) – a null hypothesis \(H_0^t\) against an alternative \(H_1^t\), \(\forall t \in D\), is provided with weak control of the FWER if the adjusted p-value \(\widetilde{p}(t)\) is such that for all \(\alpha \in (0,1)\):
\[ H_0^{D_\phantom{0}} \mbox{ is true } \Longrightarrow \mathbb{P}(\exists t \in D : \widetilde{p}(t) \le \alpha) \le \alpha. \]
Functional control of the FWER
Definition 3 (Strong control of the FWER)
A procedure for locally testing – over the domain \(D\) – a null hypothesis \(H_0^t\) against an alternative \(H_1^t\), \(\forall t \in D\), is provided with strong control of the FWER if the adjusted p-value \(\widetilde{p}(t)\) is such that:
\[ \forall A \subseteq D \text{ s.t. } H_0^A \text{ is true } \Rightarrow \mathbb{P}(\exists t \in A : \widetilde{p}(t) \le \alpha) \le \alpha, \quad \text{for all } \alpha \in (0,1). \]
Closed Testing Procedure in MDA
Definition 4: Closed testing (Marcus et al. 1976)
The closed testing procedure is a multiple testing method where a null hypothesis \(H_0^{(i)}\) is rejected at level \(\alpha\) if every intersection hypothesis that contains \(H_0^{(i)}\) is rejected at level \(\alpha\).
In terms of adjusted p-values, the closed testing procedure can be expressed as:
\[ \widetilde{p}_1 = \max \left\{ p^{(1)}, p^{(12)}, p^{(13)}, p^{(123)} \right\} \]
Extensions to functional data
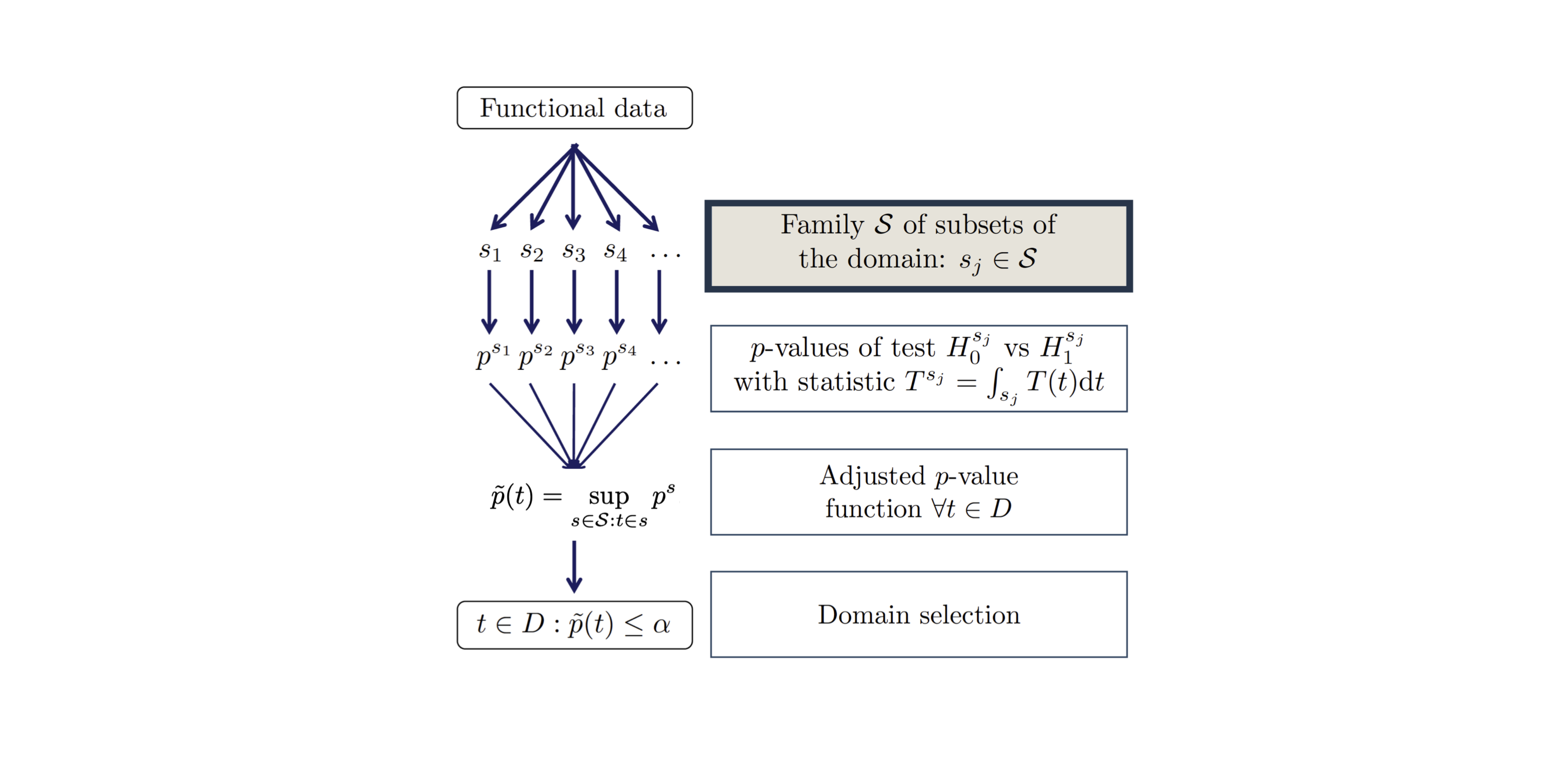
FWER-adjusted p-value function (Abramowicz et al. 2023)
\[ \widetilde{p}(t) = \sup_{A \subseteq D \, : \, t \in A} \left\{ p^A \right\} \]
FDR-adjusted p-value function (Olsen et al. 2021)
\[ \widetilde{p}(t) = \min_{s \ge p(t)} \left\{ 1, \frac{\mu(D) \cdot s}{\mu(\{r : p(r) \le s\})} \right\} \quad \text{(Benjamini-Hochberg)} \]
Unified framework for FWER control in FDA
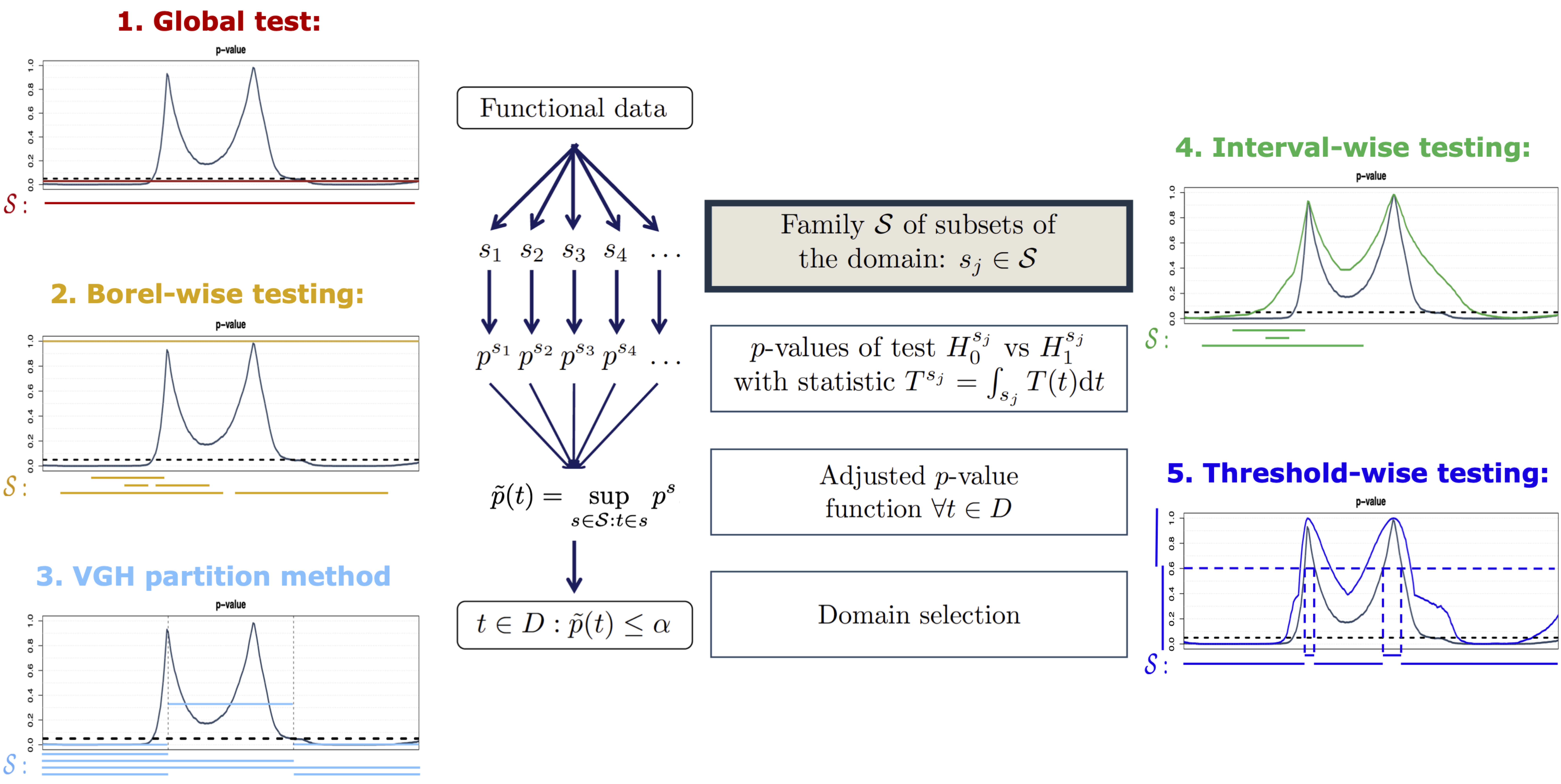
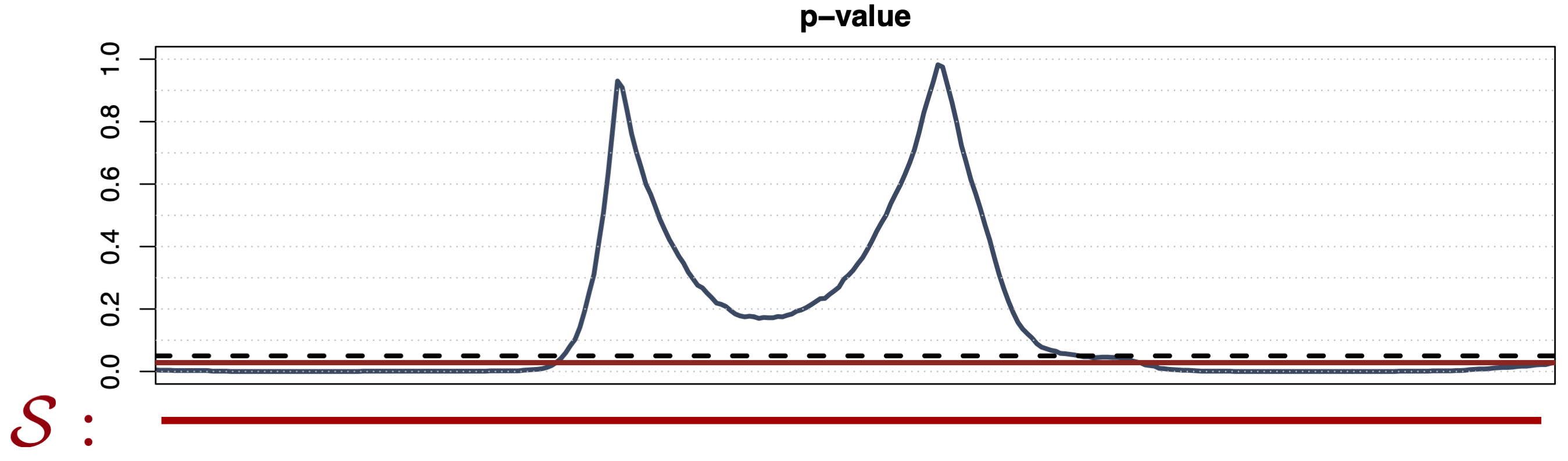
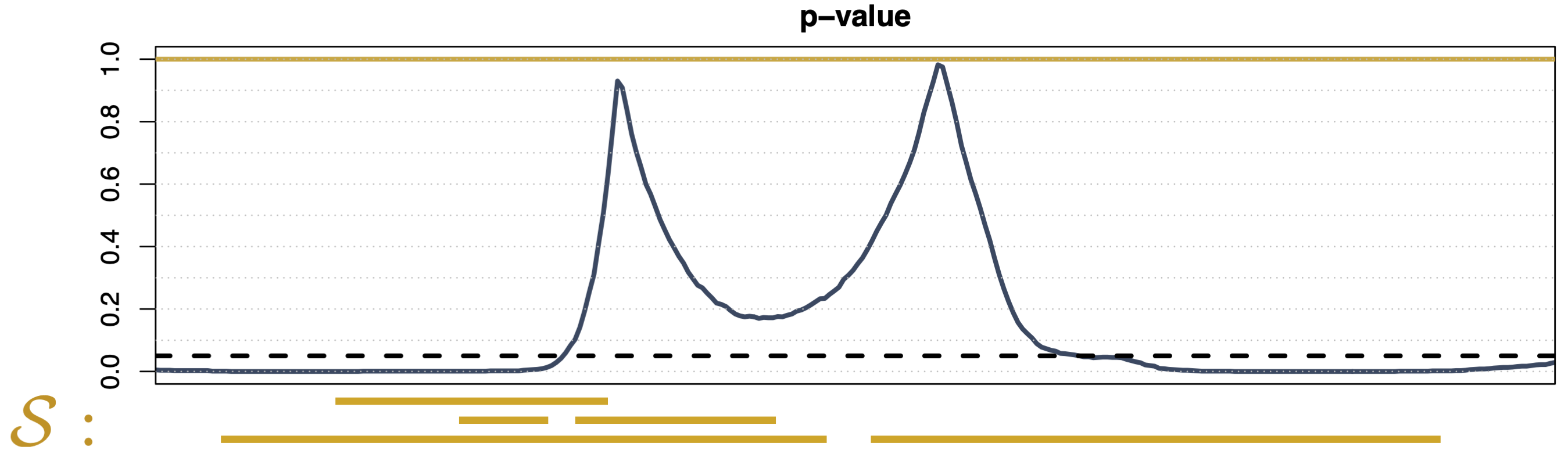
Global Testing 1
- Predefined family \(\mathcal{S} = \{D\}\)
- Weak control of the FWER
- No domain selection (i.e., \(\widetilde{p}(t)\) \(\equiv \mathrm{const}\))
- Computationally efficient (one test)
Borel-Wise Testing 1
- Predefined family \(\mathcal{S} = \mathcal{B}(D)\)
- Strong control of the FWER
- No domain selection (i.e., \(\widetilde{p}(t)\) \(\equiv \mathrm{const} \ge \max \{p(t) : t \in D\}\))
- Computationally inefficient (\(2^p\) tests for a grid of \(p\) points)
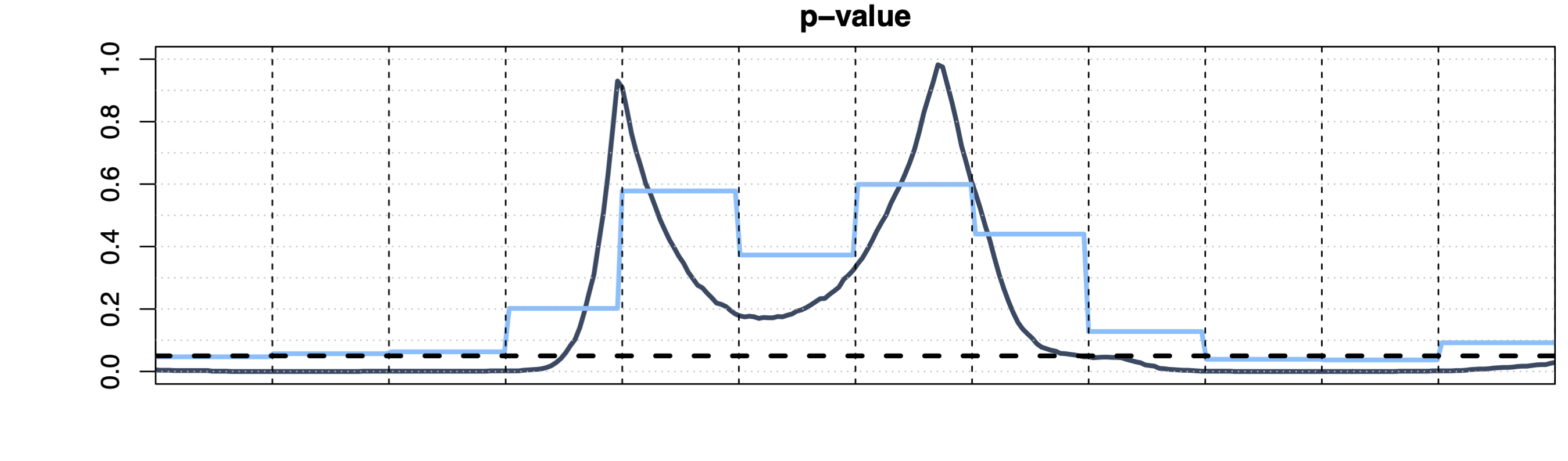
Partition-Closed Testing 1
- Predefined family \(\mathcal{S} = \sigma(\{S_j\}_{j=1}^J)\), with \(\{S_j\}_{j=1}^J\) a partition of \(D\)
- Strong between-set control, weak within-set control of the FWER
- Subset selection (i.e., \(\widetilde{p}(t)\) \(= p^{S_j}\) for \(t \in S_j\))
- Computationally inefficient if \(J\) is high (\(2^J\) tests)
Partition-Closed Testing 1
- Predefined family \(\mathcal{S} = \sigma(\{S_j\}_{j=1}^J)\), with \(\{S_j\}_{j=1}^J\) a partition of \(D\)
- For \(J\) going to infinity, PCT = BWT:
\(\lim_{J \to \infty}\) \(\widetilde{p}_J^\mathrm{PCT}(t)\) \(=\) \(\widetilde{p}^\mathrm{BWT}(t)\).
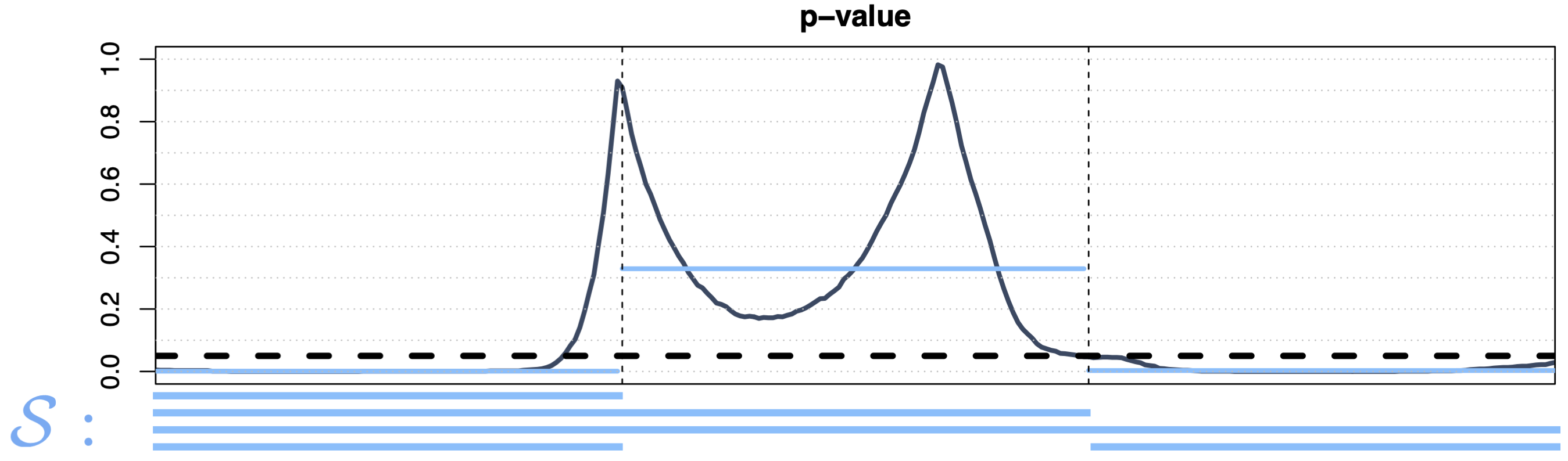
Interval-Wise Testing 1
- Predefined family \(\mathcal{S} = \{[t_1^l, t_1^r] \times \dots \times [t_d^l, t_d^r] : t_i^l \le t_i^r, i \in [\![ 1,d ]\!] \}\)
- Strong control if \(D_0\) is a Cartesian product of intervals, weak control otherwise;
- Domain selection (i.e., \(\widetilde{p}^\text{IWT}(t)\) is a continuous function)
- Computationally inefficient for large \(d\) (\(p^{2d}\) tests)
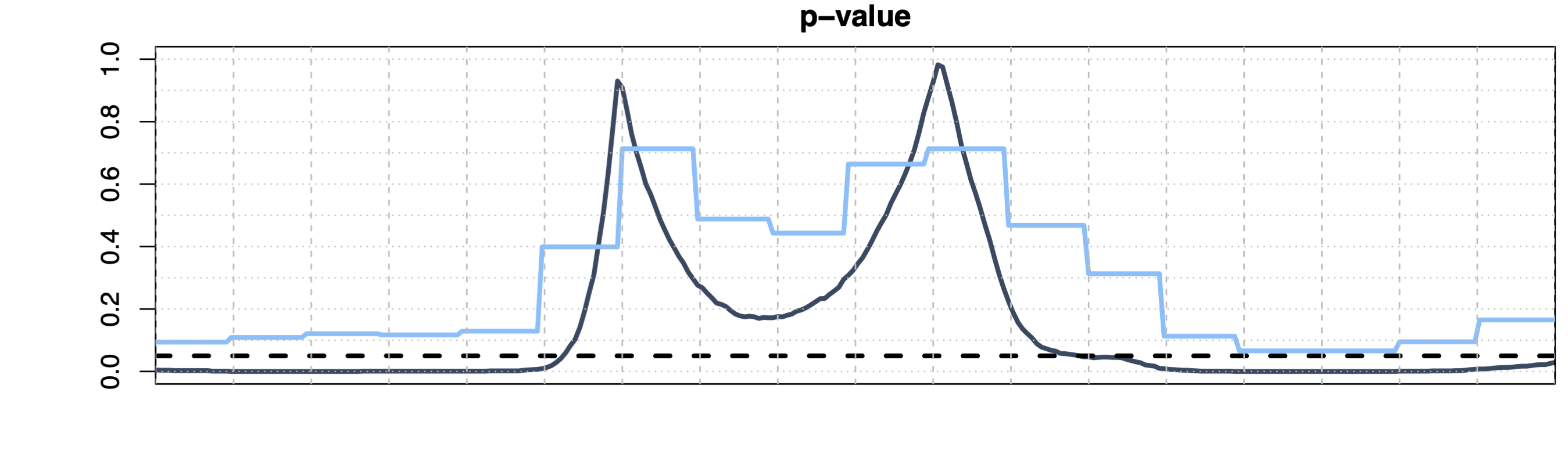
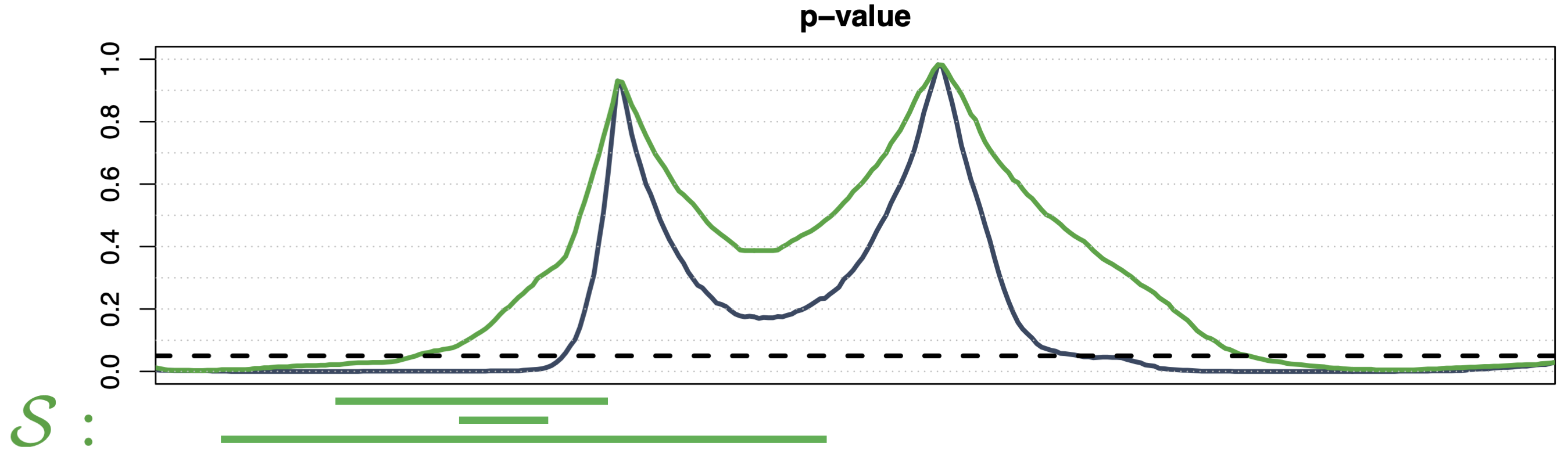
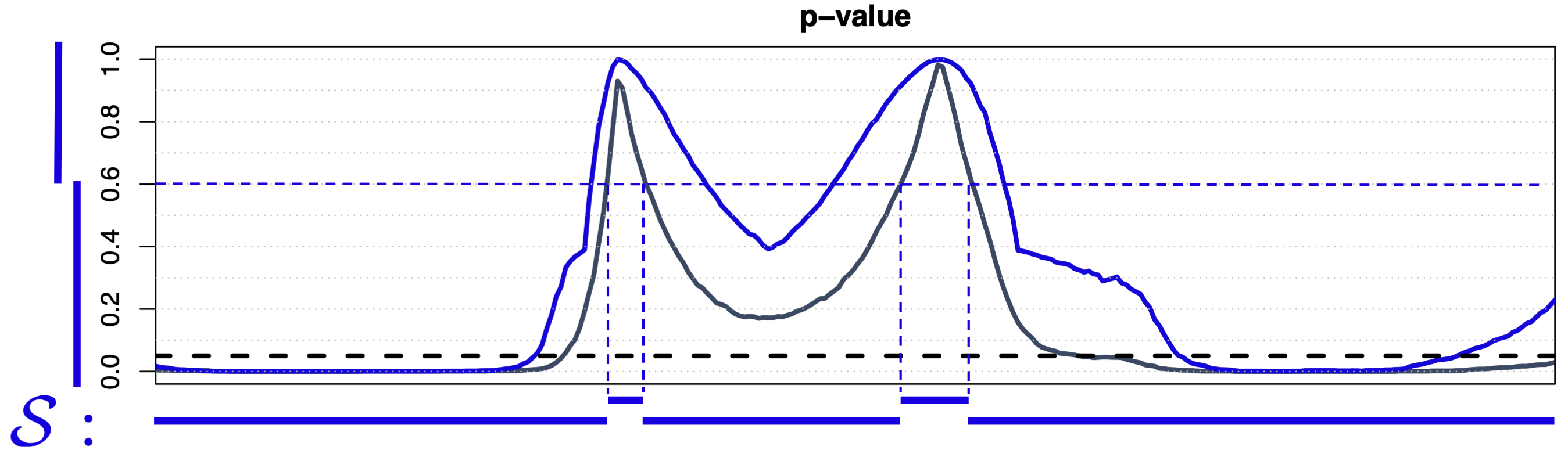
Threshold-Wise Testing 1
- Data-driven family \(\mathcal{S} = \{ \{t : p(t) \le s\}, \{t : p(t) > s\}, s \in [0,1] \}\)
- Strong control asymptotically, finite weak control
- Domain selection (i.e., \(\widetilde{p}^\text{TWT}(t)\) is a continuous function)
- Computationally efficient for any \(d\) (\(2m\) tests, with \(m\) the discretization step of \([0,1]\))
A more complex example
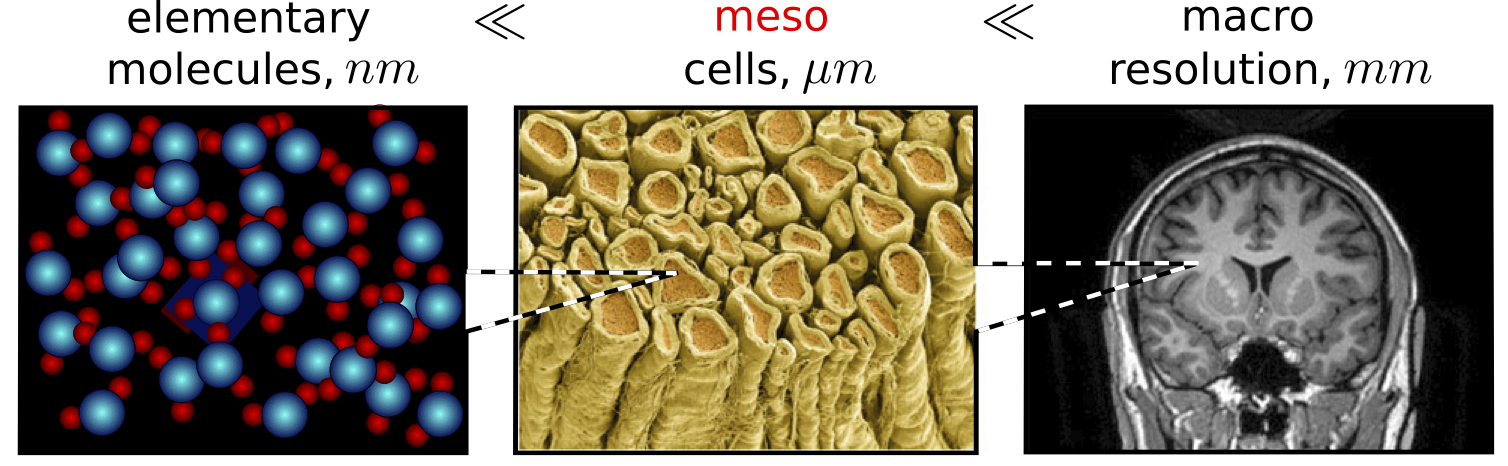
Brain microstructure mapping
- Diffusion MRI: sensitive to diffusion of water molecules in the brain;
- Diffusion model: locally describes parametrically the diffusion pattern.
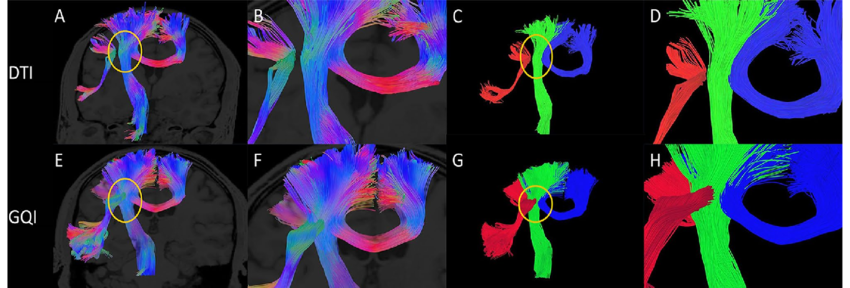
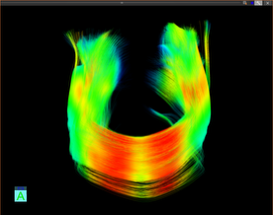
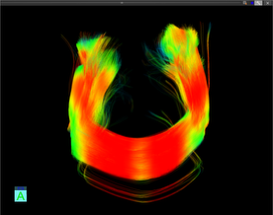
Brain structural connectivity mapping
- Pyramidal tract (green): motor function
- Arcuate fasciculus (red): language function
- Corpus callosum (blue): interhemispheric communication
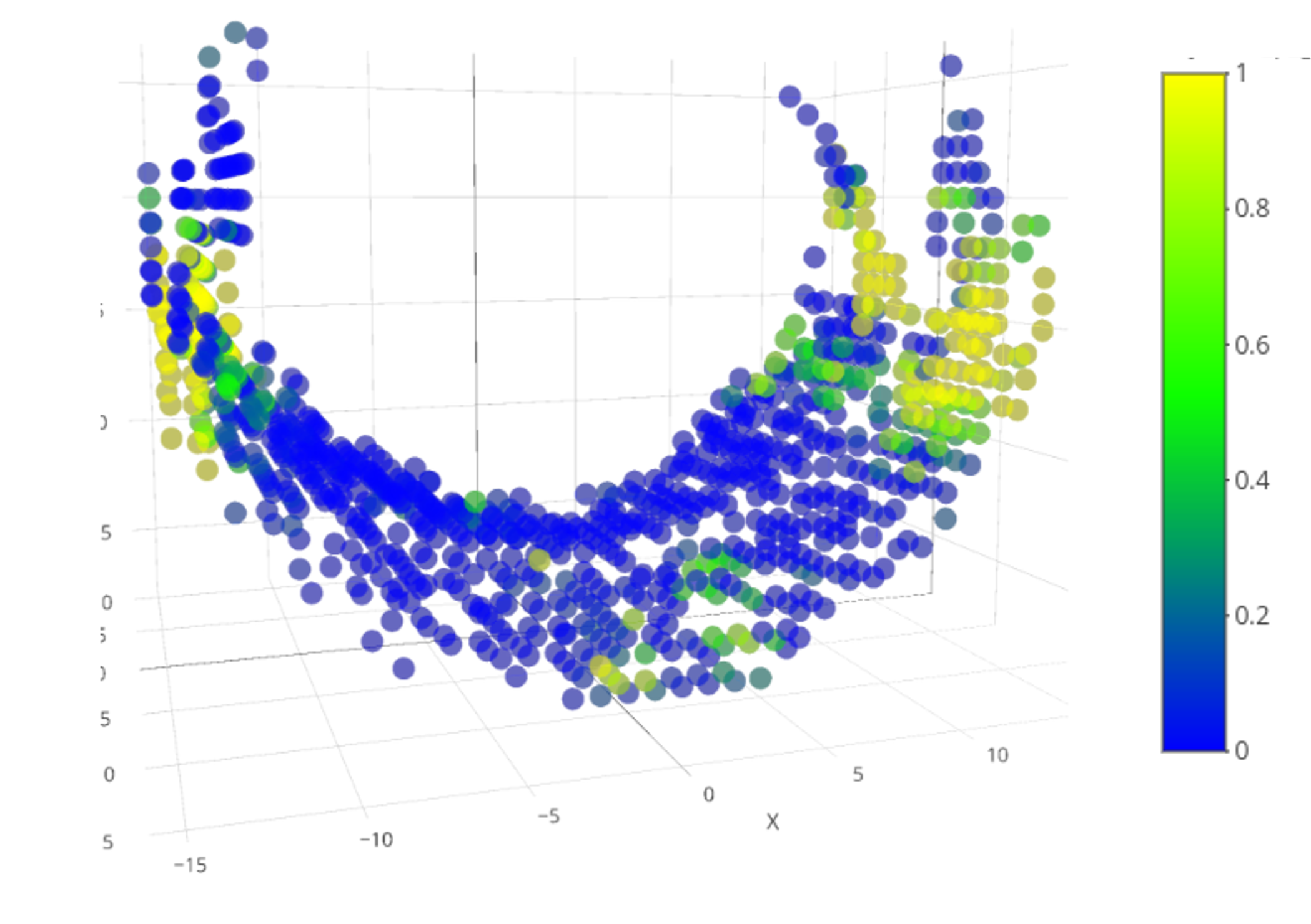
Conducted study
- Lesions = axonal damage \(\Rightarrow\) drop in fractional anisotropy (FA).
- Study the FA of healthy subjects along the corpus callosum (CC).
- Compare FA stability along CC when two diffusion models are applied:
Results
- We applied TWT to test differences in FA variance between M1 and M2;
- We found significantly lower FA variance when using M2 compared to M1;
- Non-significant regions correspond to areas of the CC that cross with pyramidal tract and arcuate fasciculus \(\Rightarrow\) both models are misspecified in these regions.
Software
Implementation: the R package {fdatest}
Achieved
Initial release following IWT publication (Pini and Vantini 2017).
Implements IWT for:
- One-sample problems;
- Two-sample problems;
- Functional ANOVA;
- Functional regression.
Compatible with the {fda} package.
Implementation: the R package {fdatest}
Todo
Almost done in branch astamm/fdatest@clean-doc, which you can try out by installing the package from GitHub: